PCA of certain columns R1/14/2024  
We also document contemporary practices by a literature survey of 125 representative articles that apply PCA to SNP data, find that virtually none implement our recommendations. 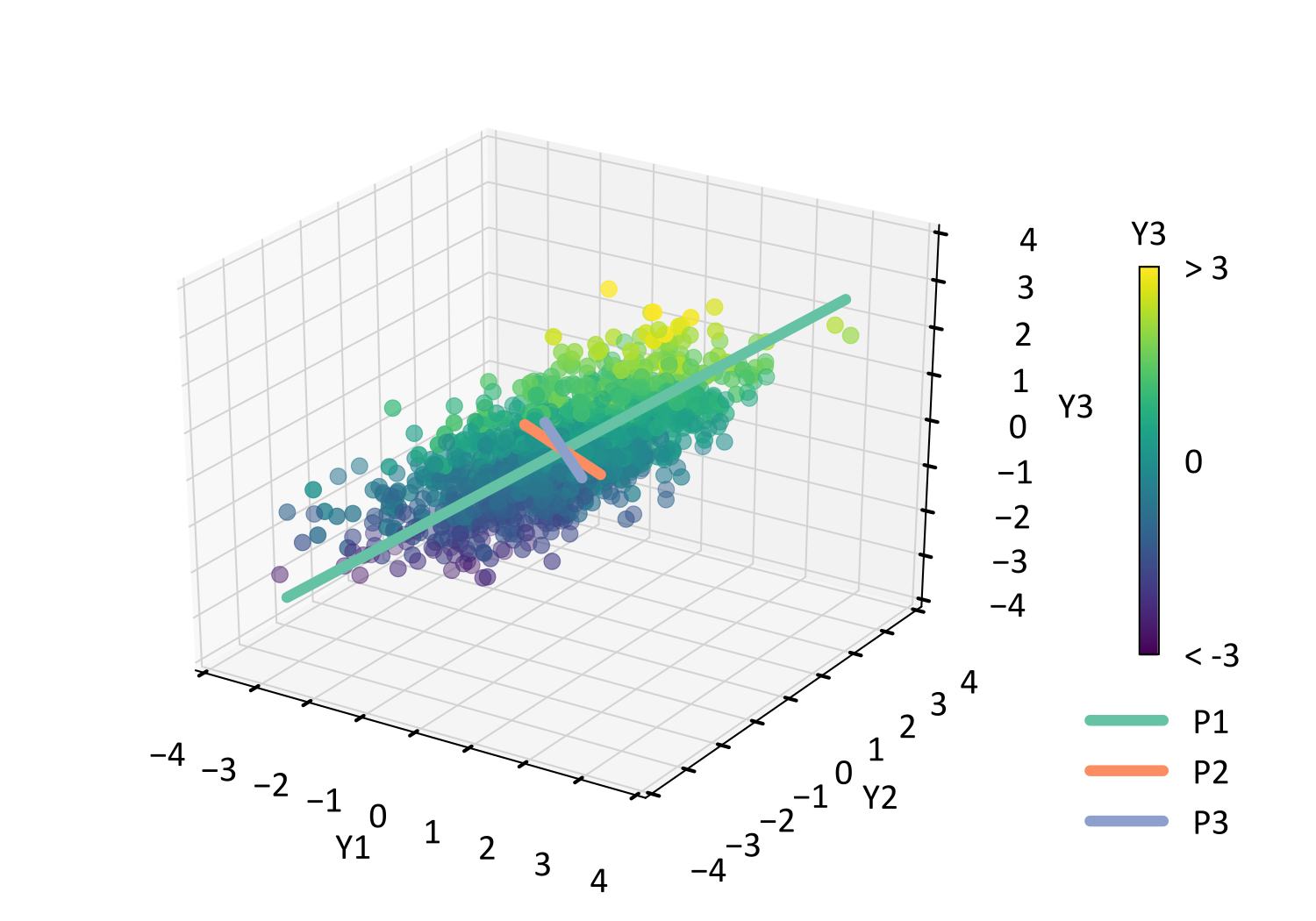
Our main three recommendations are simple and easily implemented: Use PCA biplots, SNP coding 1 for the rare allele and 0 for the common allele, and double-centered PCA (or AMMI1 if main effects are also of interest). PCA is not a single method that is always done the same way, but rather requires three choices which we explore as a three-way factorial: two kinds of PCA graphs by three SNP codings by six PCA variants. Accordingly, PCA graphs are frequently used to provide a low-dimensional visualization in order to display and discover patterns in SNP data from humans, animals, plants, and microbes-especially to elucidate population structure. 
SNP datasets are high-dimensional, often with thousands to millions of SNPs and hundreds to thousands of samples or individuals. 
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
